HGNC informatics
Web statistics, releases and development
Kristian Gray
Contents
- genenames.org releases
- Gene families
- Search
- New BioMart server
- GitHub: code release
- Web statistics
- New internal curator tools
- Xref updater
- New mapping tool
genenames.org releases
Gene families
- Released new gene families resource (beta Jan, live Apr).
- Gene families are discussed in detail in Ruth's talk.
- How successful was the release? (See web statistics section).
- Ruth and myself have written a Gene families paper (to be submitted to Human Genomics).
Search
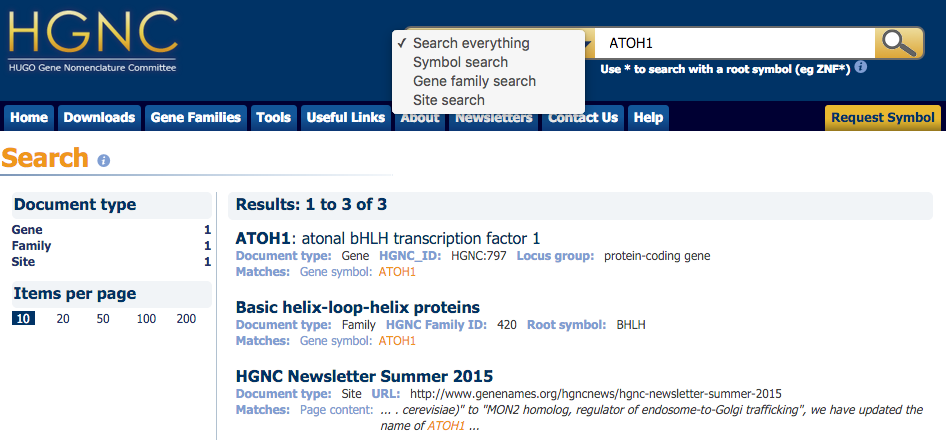
Can now search all aspects of the site from one search form.
GitHub: code release
Several javascript visualisation tools were created whist creating the gene family resource which could be useful to other developers. Therefore I am releasing the code as open source on github. So far I have release:
euro-pmcentralizer
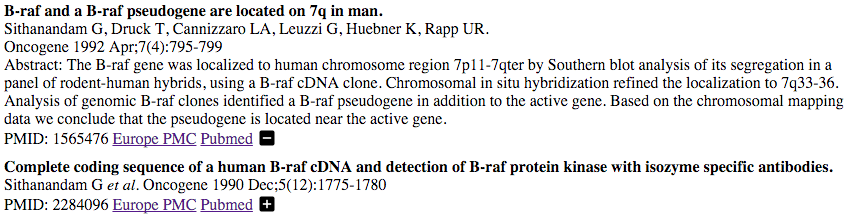

pfam-dom-draw

Old BioMart server
- Old server was to be decommissioned.
- We didn't have admin rights to make changes easily.
- Some of the functionality of the old BioMart service no longer worked.
- Decommissioning offered an opportunity to create a new service and add more functionality.
New BioMart server
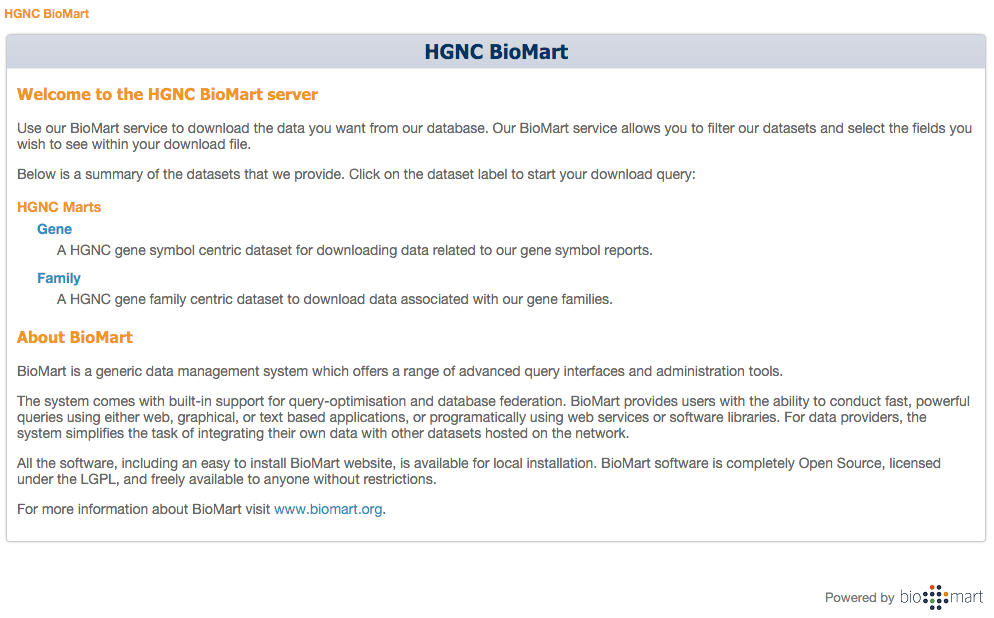
The BioMart home page offers the user a choice to search and download gene or family centric data.
Family mart
New mart service to join the gene mart portal to search and download family centric data. Users start by adding query terms to the filters section...
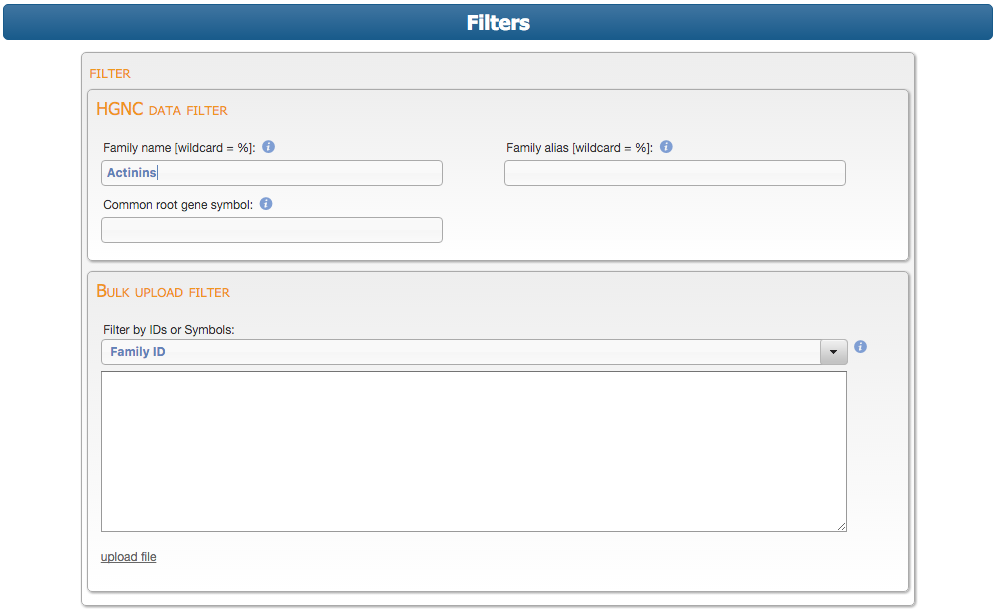
...then select which attributes they would like to see in the results table.
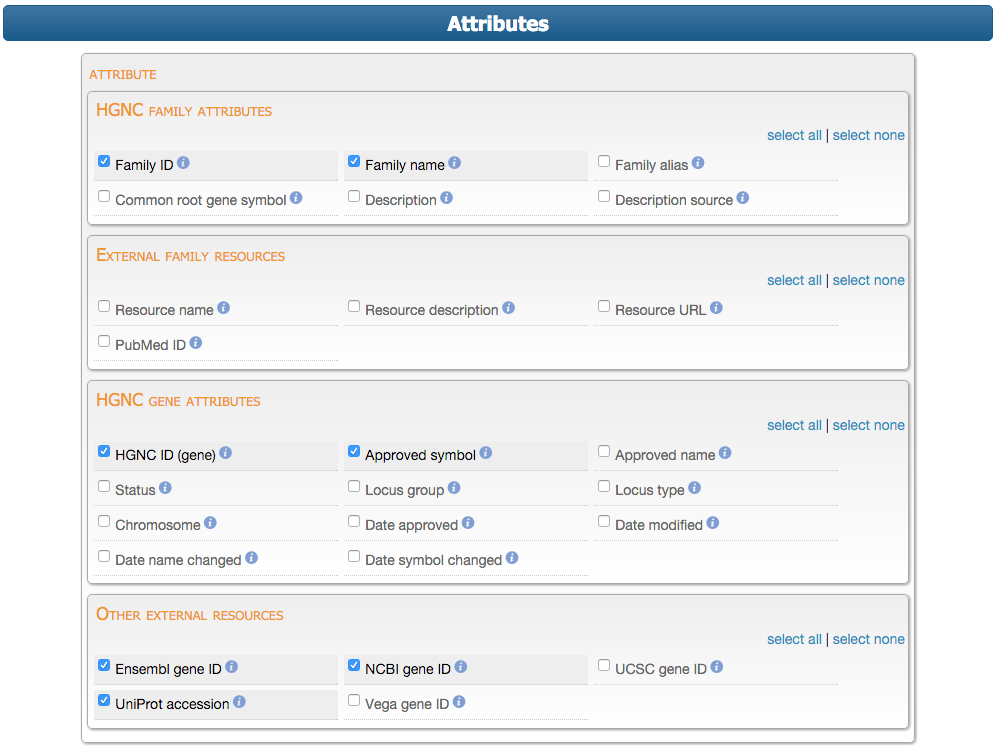
Submitting the form then produces a preview results table and the option to download the data is presented.
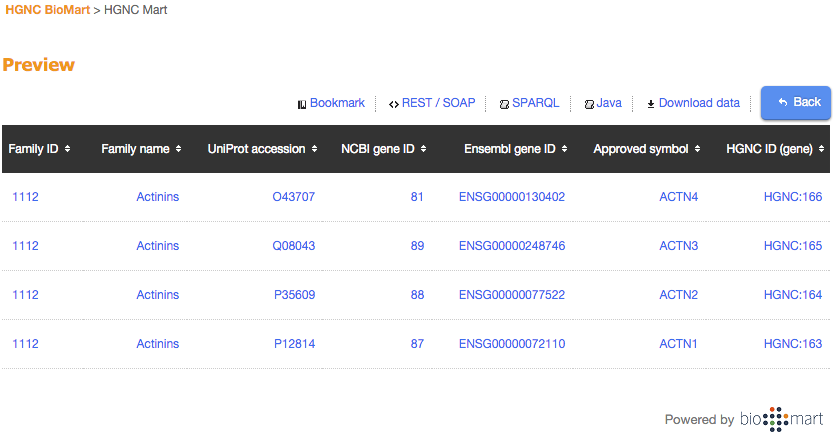
Web statistics
Number of users
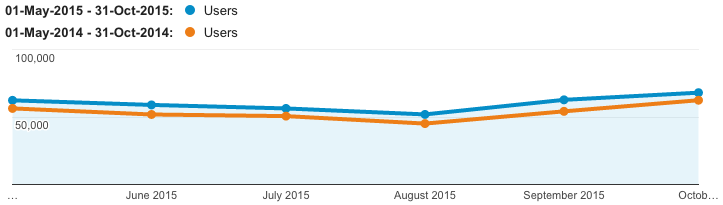
| Total users | Mean | |
|---|---|---|
| 2015 | 327,635 | 56,000 per month |
| 2014 | 291,737 | 53,435 |
↑12% over the 6 month period
User behaviour
Oct 2015 compared to Oct 2014| Date | Total page views | Resource | page views | Percentage of page views |
|---|---|---|---|---|
| Oct 2014 | 247,043 | Gene symbol reports | 118,094 | 48% |
| Old gene families | 12,679 | 5% | ||
| Oct 2015 | 262,586 ↑6% | Gene symbol reports | 127,275 | 45% |
| Old gene families | 5886 | 2% | ||
| New gene families | 18,955 | 9% |
Figures show that we have an increased number of page views for gene families after releasing our new family resource.
Who are our biggest referrers?
Huge increase in people coming to our site directly.
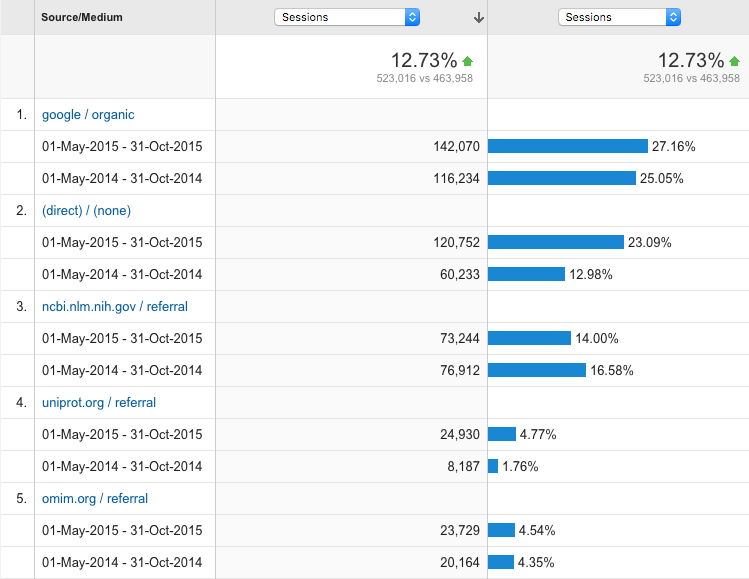
Most popular cross reference links
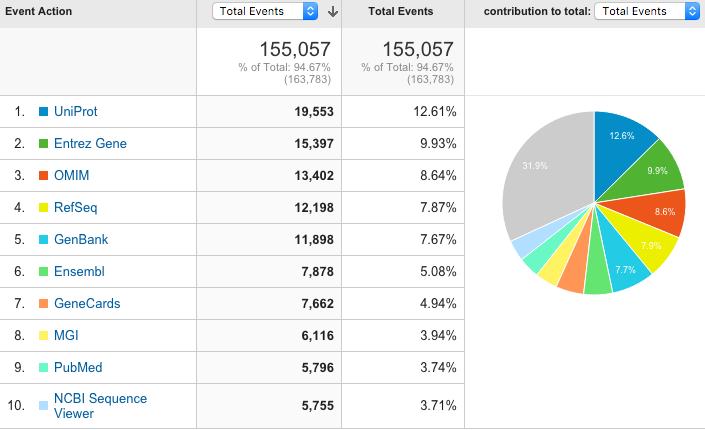
The table is similar to last year with the UniProt links being the most popular. Ensembl links have become more popular compared to last year, rising from 8th to 6th in the table.
Locations
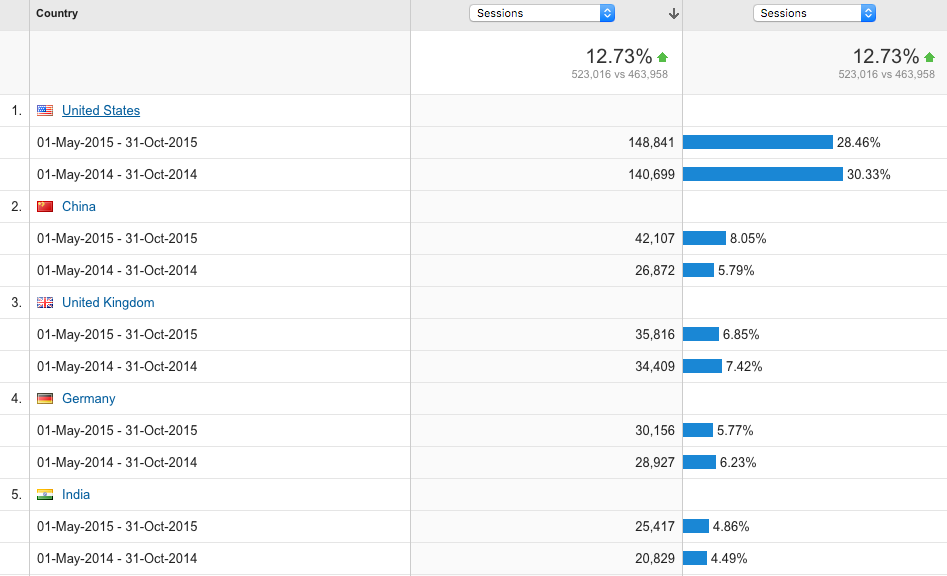
US is by far the biggest user of our resources, however the number of sessions from China has increased by 57%.
Internal curator tools
Xref updater
New set of tools to help the curators to update and add cross references to other databases. All the Xref updater tools use the HGNC ID to link our resource to others.
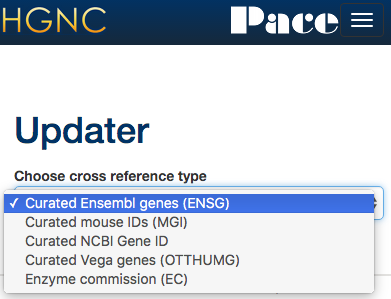
Xref updater - NCBI gene
Uses data downloaded from NCBI and compares the NCBIs gene IDs.
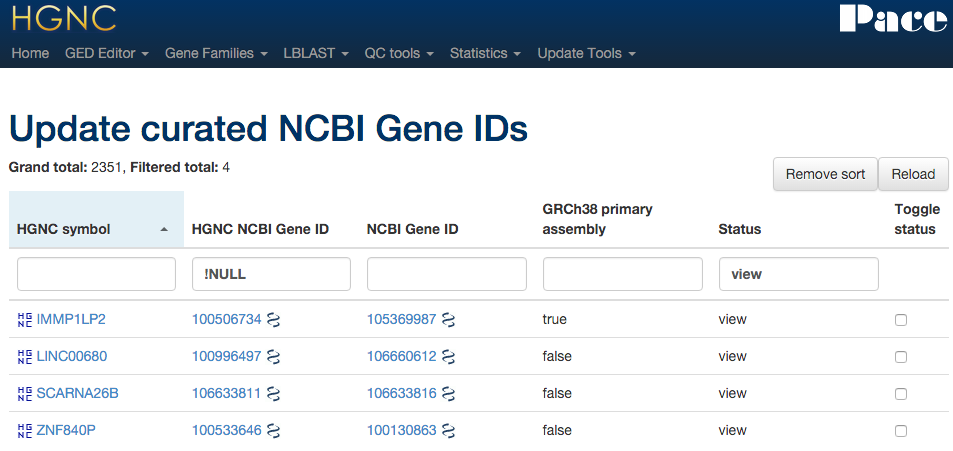
Xref updater - Ensembl
Directly connects to the Ensembl DB to compare our ENSG ID with Ensembl's.
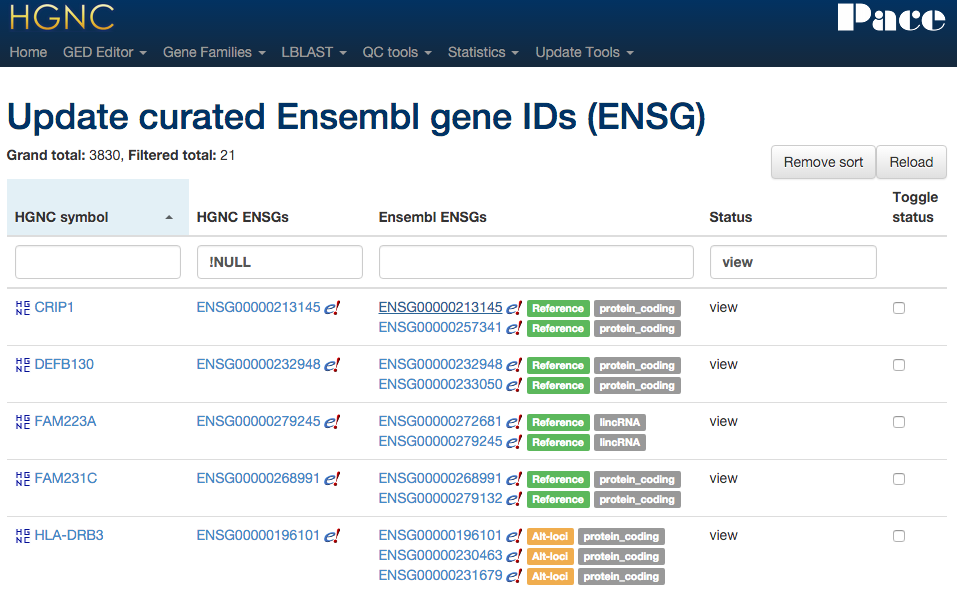
Xref updater - Vega
Connects to the Ensembl & Vega DBs directly to compare OTTHUMG IDs via HGNC ID in all three resources.
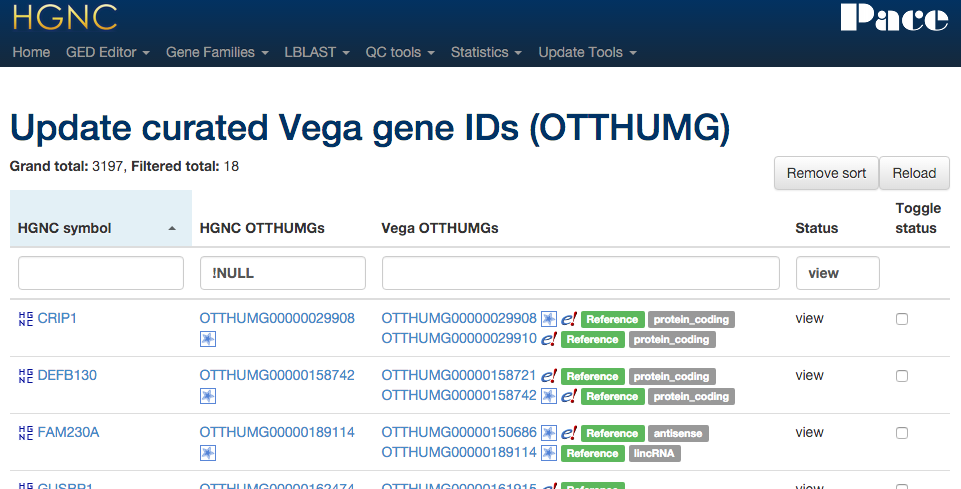
Mapping tool
The previous xref updater tools reports mismatches between our resource and the others. This new mapping tool edits the cross references per gene, and provides ensembl like tracks to compare NCBI gene, Ensembl and Vega on the GRCh38 build.
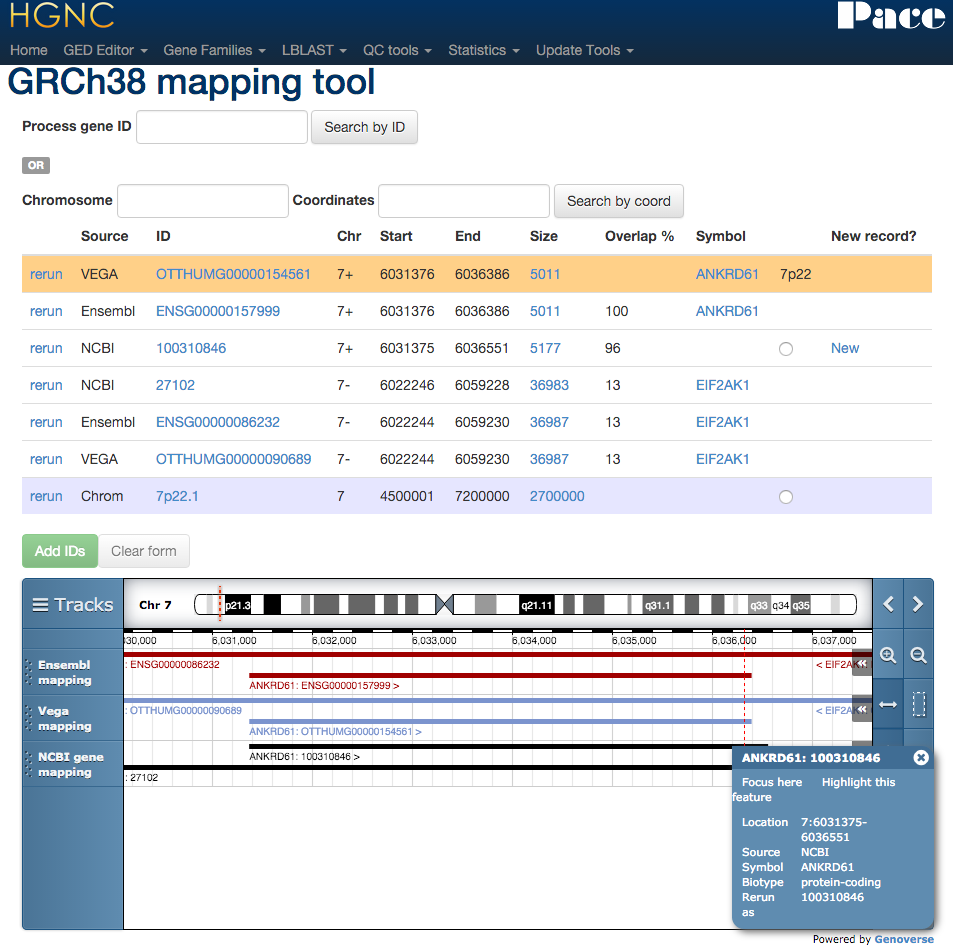
Xref updater - Enzyme commission
Compares our EC numbers with the ones found in UniProt via HGNC ID.
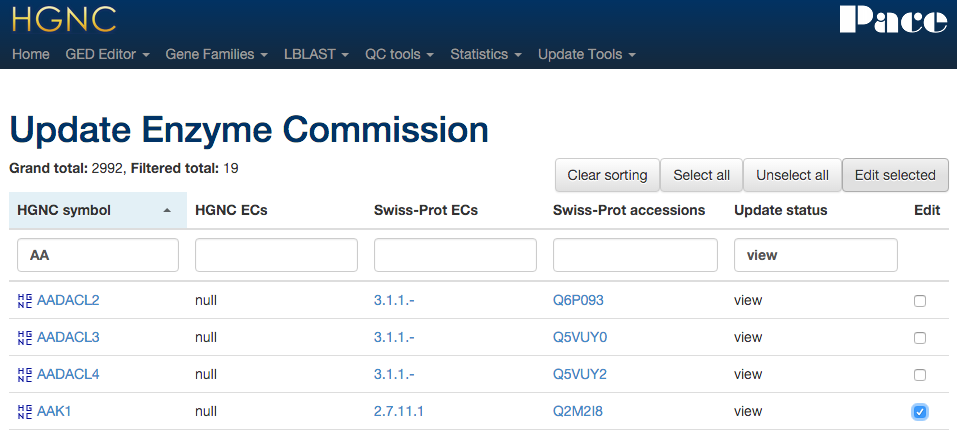
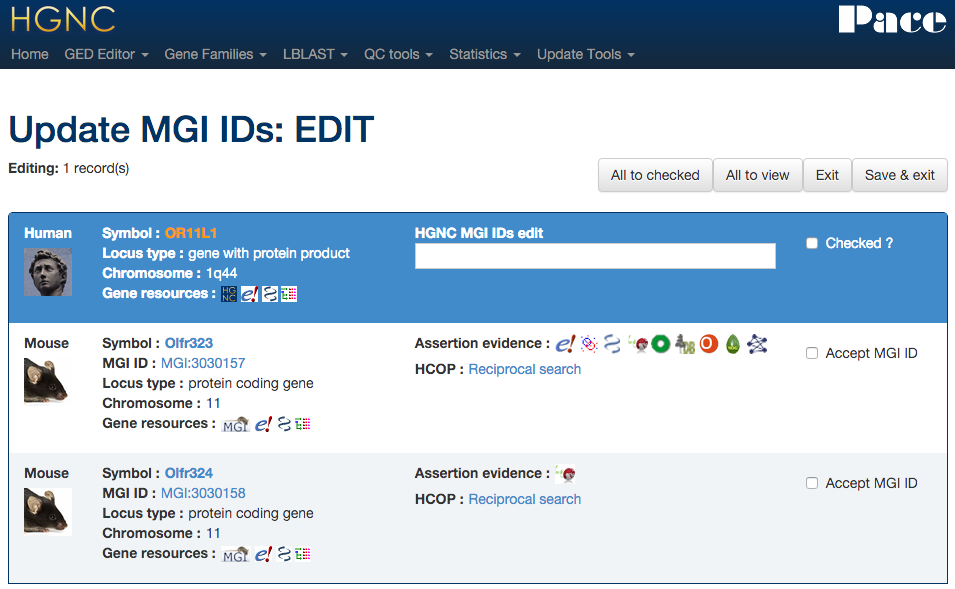
Xref updater - MGI
Using the data found in HCOP we can compare and assign MGI mouse ortholog IDs. Selecting rows to edit and pressing "Edit selection" ...
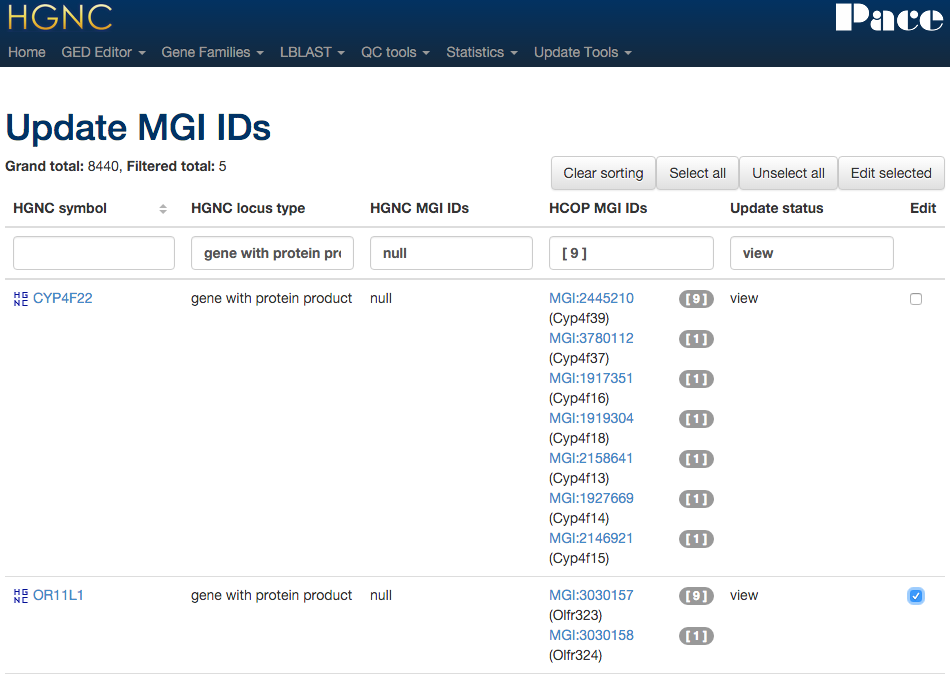
Xref updater - MGI
... will take the curator to a form resembling a HCOP results page where the curator can select the MGI ID(s) to link to the HGNC gene record. Like the EC updater, multiple genes can be edited at once in one form.