Website & database
Web statistics, releases and development
Kristian Gray
Contents
- Hardware and infrastructure changes
- Released changes & additions
- Home page
- Search
- Gene symbol reports
- REST service
- Other improvements
- Web statistics
Hardware & infrastructure
Previous infrastructure
- Originally two servers (dev & live) here at Hinxton
- The team could only control the dev server
- Releases had to be conducted by the web development team via email
- Servers were on old hardware which our systems team wanted to decommision
Slow development + no fail over + aging hardware
= high risk of lengthy denial of service
New external web architecture
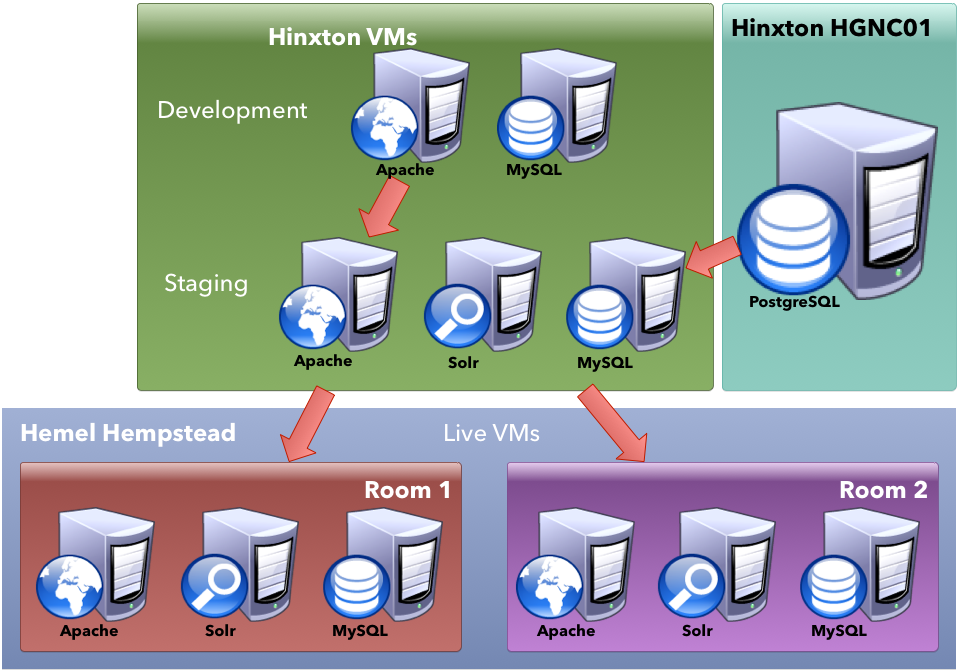
Recent releases
Home page
- Text on the page is clearer to read and more concise
- New functional word cloud which activates a search for common root symbols
- New search bar within our masthead
Original HGNC Search
- HGNC "Quick Search" was totally made in house
- Suited the users needs
- However ... problems with scalability
- Difficult to maintain and reuse for other purposes
- Well known search platform used by many companies and widely used across campus
- Highly scalable & very quick
- Easy to maintain & separate from our code base
- Provides faceting and highlighting out of the box
- Can use wildcards, phrases and logical operators
- Can limit the search to one field
Gene symbol reports
- Our most important pages
- Slightly different layout and added some improvements
- New functionality: help information and references
REST service
- Introduce a new REST service to retrieve data from within our database
- Built upon our Solr search server
- Users can return data within an XML or JSON format for easy parsing
- Three main commands:
- info
- search
- fetch
Other improvements
- Updated list search and renamed it "Multi-symbol checker"
- A tool for our users to check a multiple gene symbols
- Updated the interface
- New code to make the search quicker for large lists
- Downloads & statistics page
- Added statistics for genes which reside in alternative loci only
- Karyotype image controls the statistics per chromosome
- Data sets are no longer created on-the-fly and are stored within the EBI FTP site
Web statistics
All statistics were collected by Google analytics between
May 1, 2014 - Oct 28, 2014
Number of users
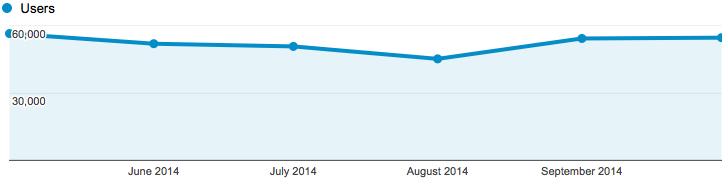
- Mean = 47,000
- Median = 53,000
Where do the users come from?
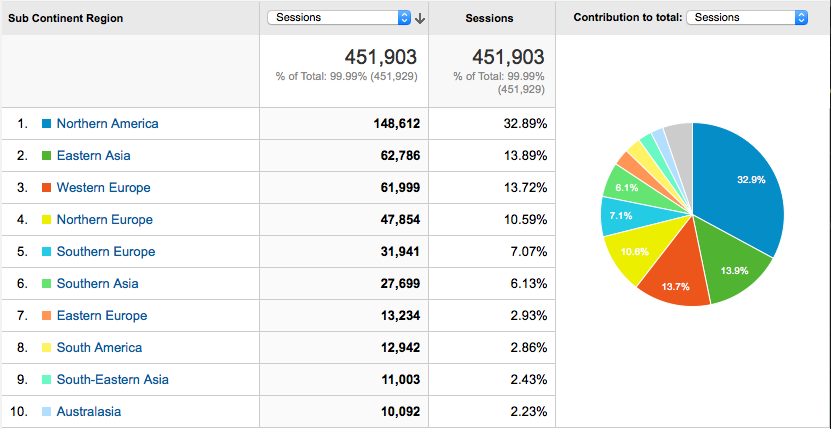
User behaviour
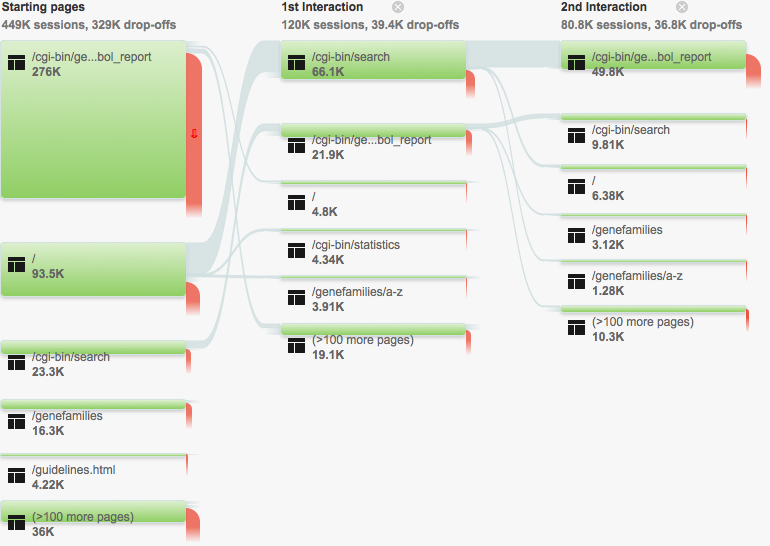
Who are our biggest referers?
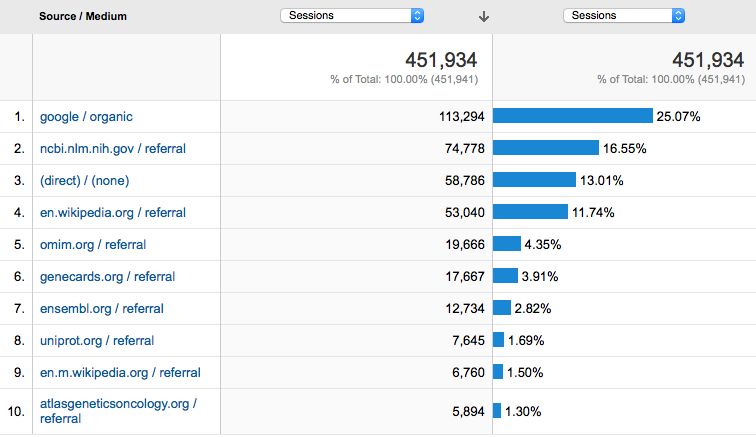
Where do our users go?
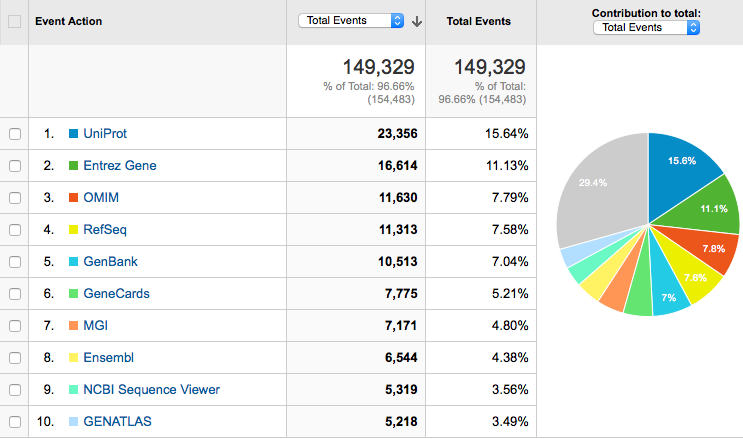
In summary...
- We are widely used
- Steady number of users of 53,000 users per month
- Important role of linking resources together as well as providing gene nomenclature
- We are as widely used within North America (33%) as we are in Europe (34%).
- Asia (23%) an emerging user base
