What's going on in the HGNC?
A whislestop tour of what we are working on now and in the future
Kristian Gray
What do the HGNC do?
HGNC aims
- To create and approve nomenclature for all human protein-coding genes, pseudogenes & RNAs.
- Reassign nomenclature for loci basing names on new functional data.
- Group genes into families/sets.
- Coordinate gene naming across vertebrates.
- Collaborate with other nomenclature committees e.g Mouse, Rat and Chicken.
- For vertebrates without a nomenclature committee we have created a new project called VGNC.
Specialist advisors & collaborator
- Specialist advisors
- 116 people across the world that are our specialist advisors for particular gene families.
- Full list can be seen on our specialist advisors page
- Collaborators on complex gene families
-
- Naming and grouping olfactory receptors
- Tsviya Olender & Doron Lancet
- Naming and grouping P450 genes
- David Nelson & Jed Goldstone
Naming and curating genes
Assignment of gene symbols
- Function
- e.g. enzyme, complex subunit, receptor, transporter
- Sequence comparison
- e.g. ortholog, gene family member
- Domain structures & motifs
- e.g. TMEM#, WDR#
- Other information from researchers, literature & databases
- e.g. cellular location, associated phenotype, chromosomal location
Reassignment & coordination
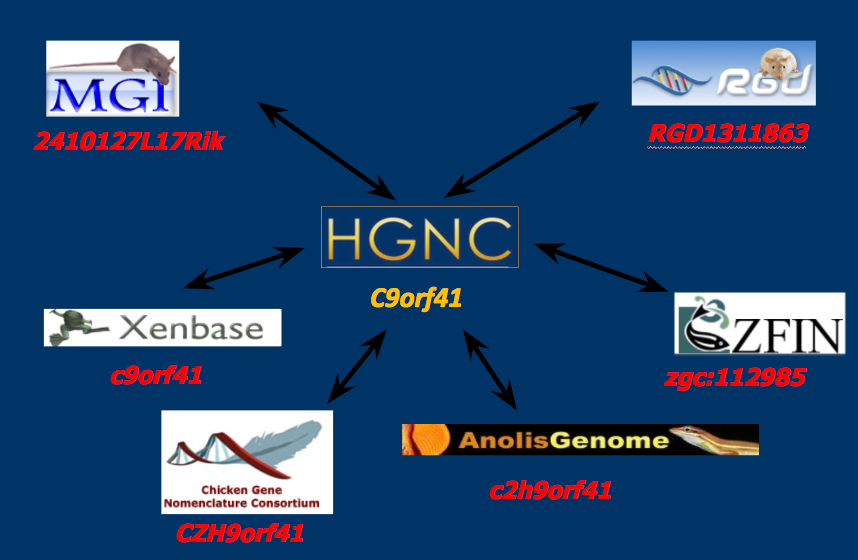
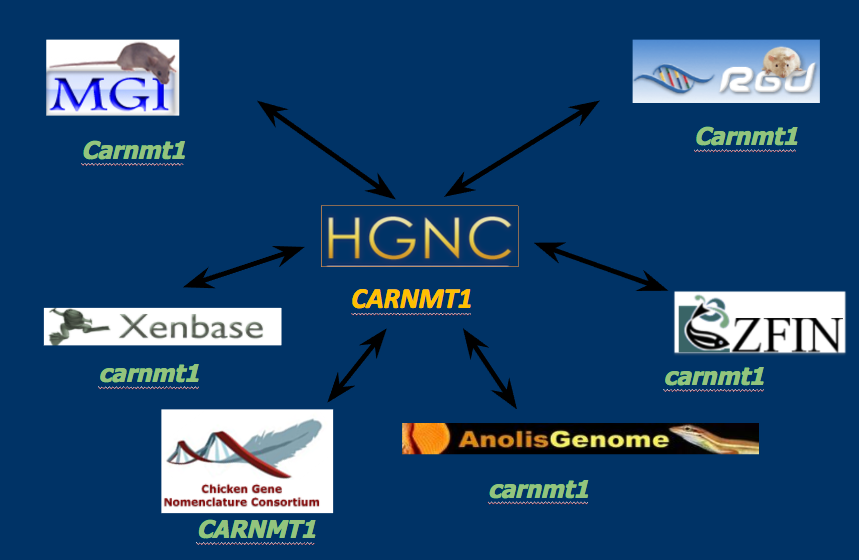
Tools & resources

Curated cross references
Let's say that we have created a gene symbol record using NCBI gene 5989 and named it RFX1.
We can use the NCBI gene ID that we have attached to the record to curate links to Ensembl, Vega* and pseudogene.org*
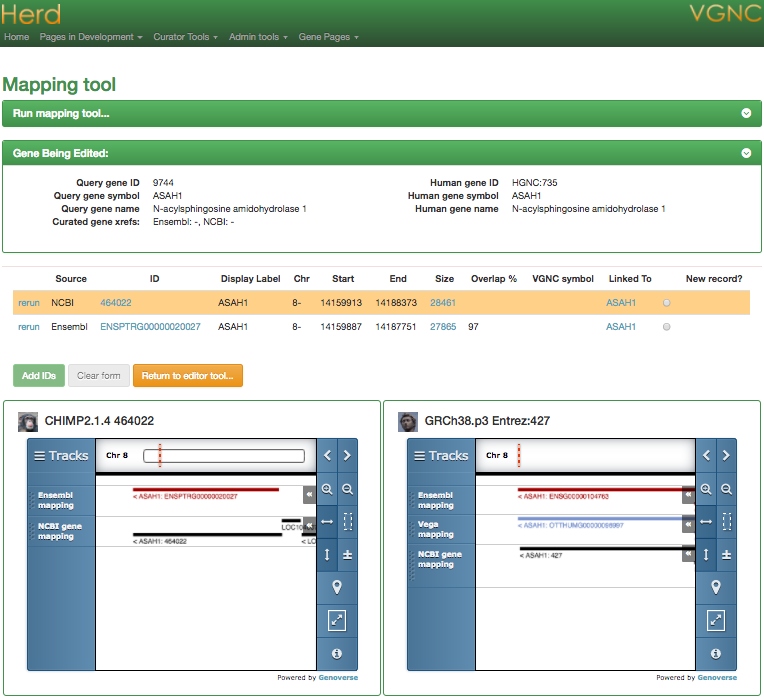
Mapping Tool
Maintaining x-refs
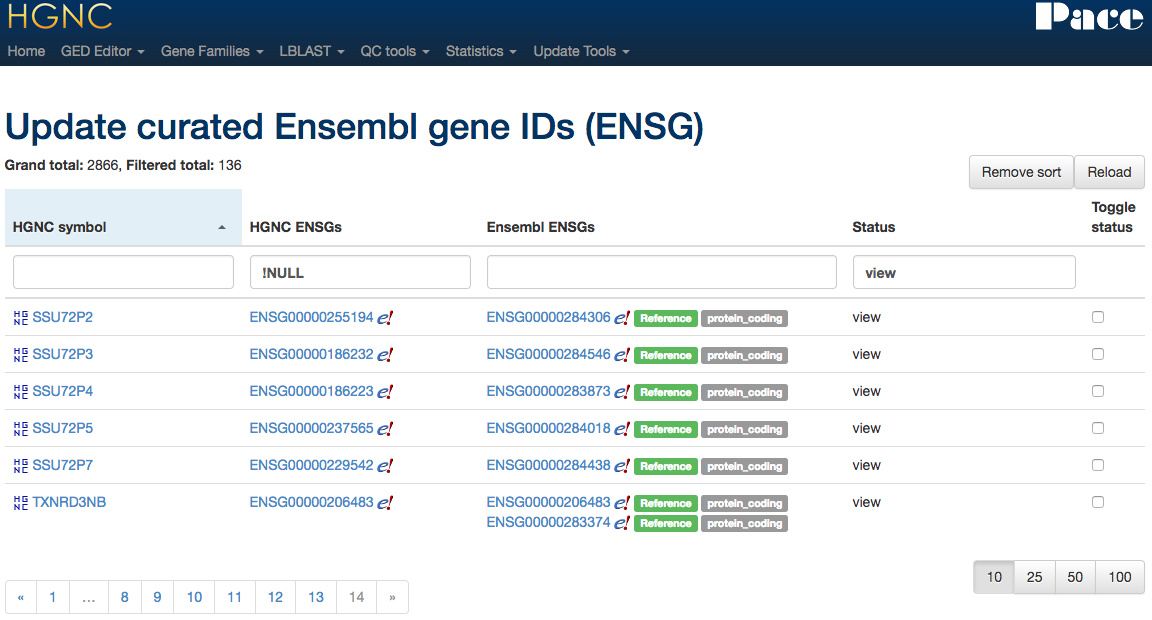
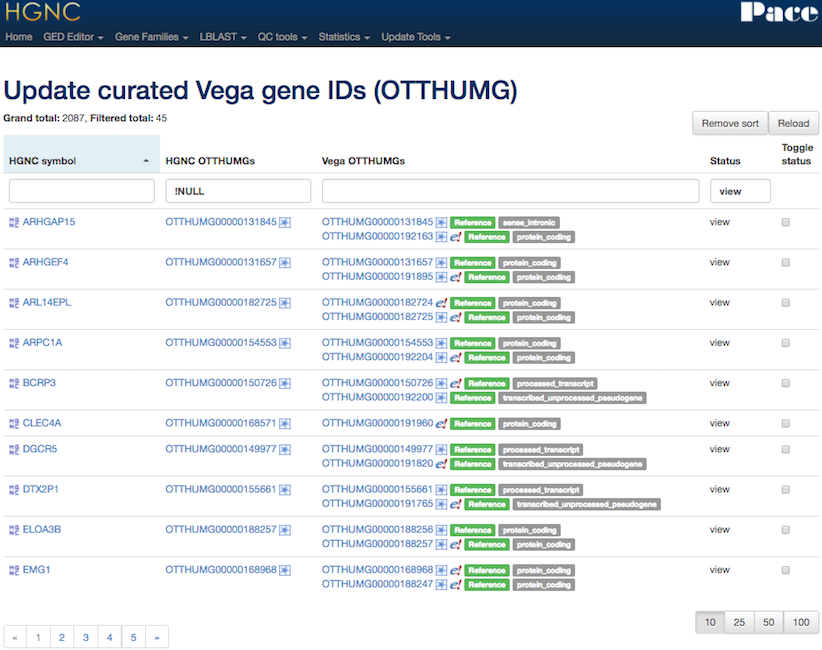
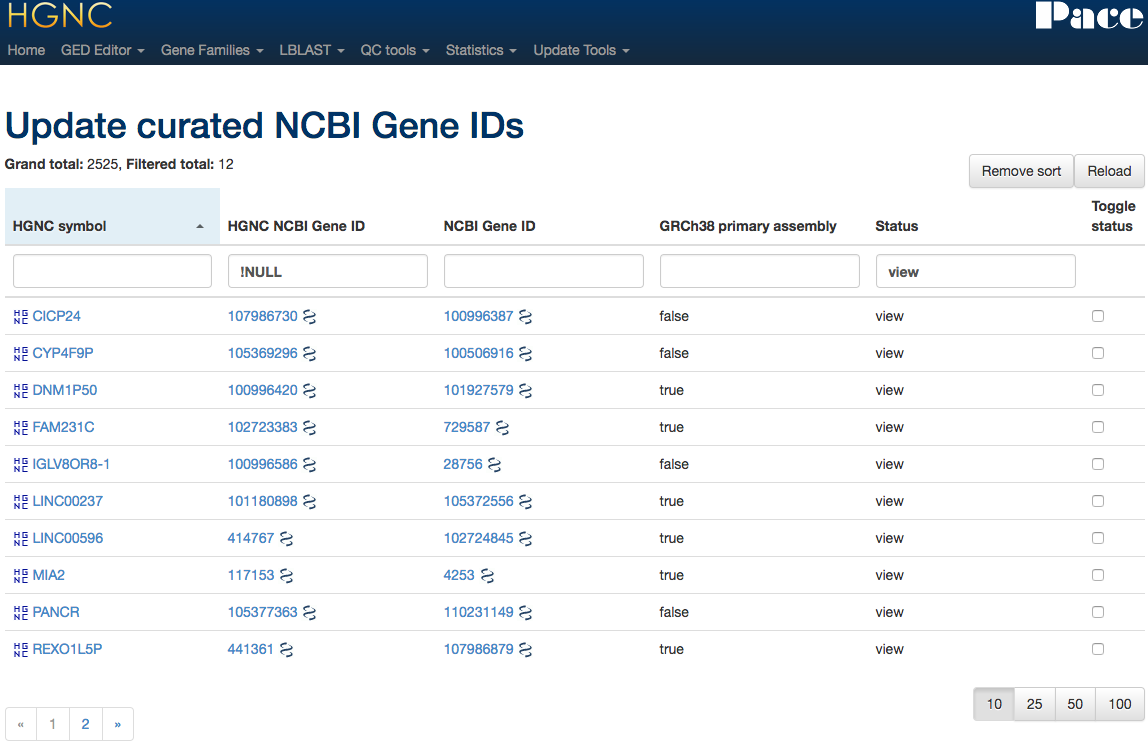
Gene family curation
Actin familyNew HGNC website development
The current HGNC site
- Static content served by Drupal 6 while dynamic content is perl CGI. This means two live servers with two sets of code and templates.
- Site is fixed width for smaller monitors.
- Not mobile or tablet friendly.

Beta website
- Worked with the UX team to see what we could do to improve the site.
- Cleaner design.
- Familiar flow through the website.
- Improved search and facet options.
- All running off one web server using AngularJS (1.6) and Jekyll.
- Development carried out on developers own machines within a docker container cluster.
- ... and managed by the Gulp task runner to build, uglify and compress before release.
- Carried out UX testing.

Planning on releasing the beta site to everyone this year.
VGNC
Vertebrate gene nomenclature committee
What is the VGNC?
- New initiative to coordinate gene naming across vertebrates
- Gene nomenclature will reflect homologous relationships across species
- Consensus naming based on human
- Data will be distributed to genomic databases and model organism databases
- The new VGNC website will become a portal for all official vertebrate nomenclature
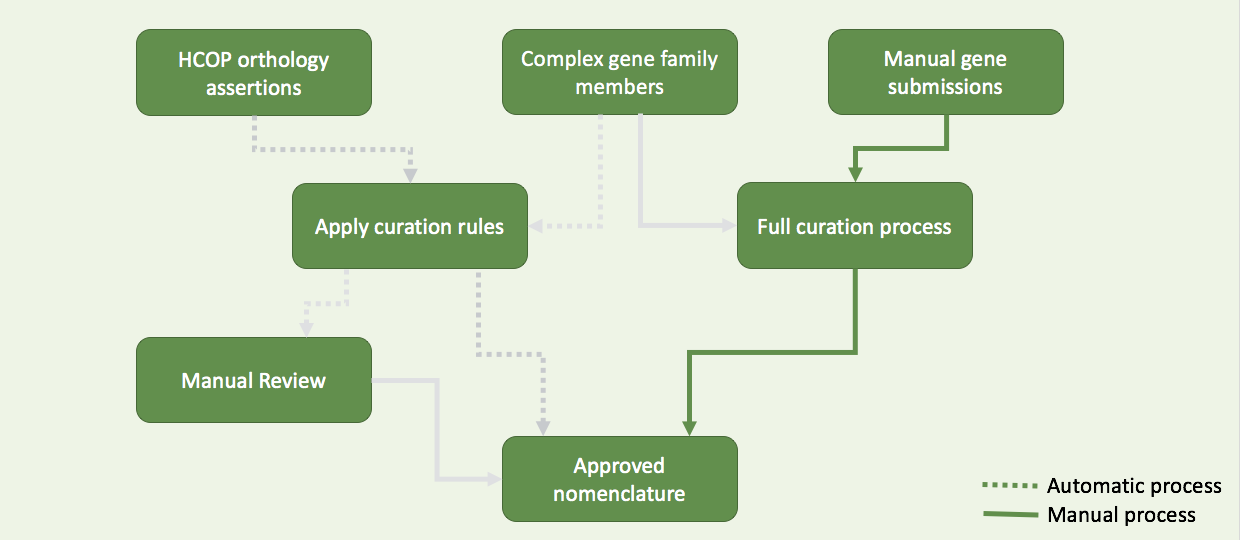
HCOP orthology assertion
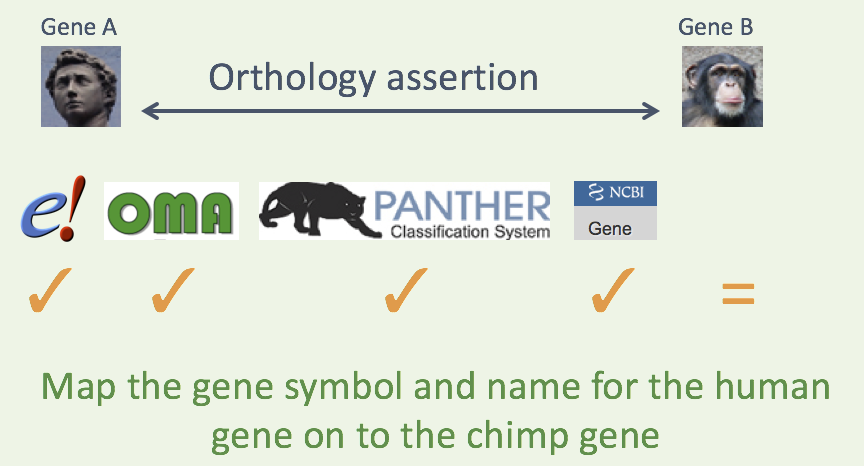
Curation tools
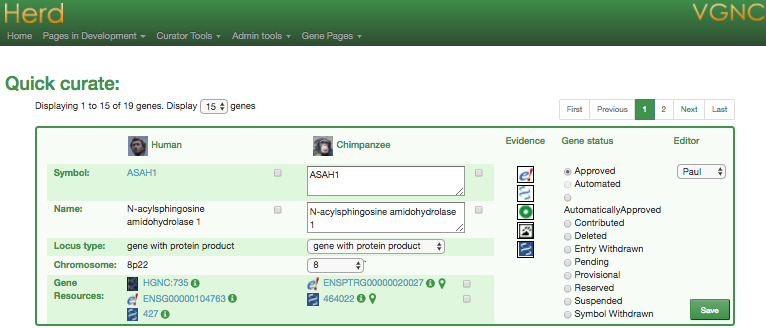
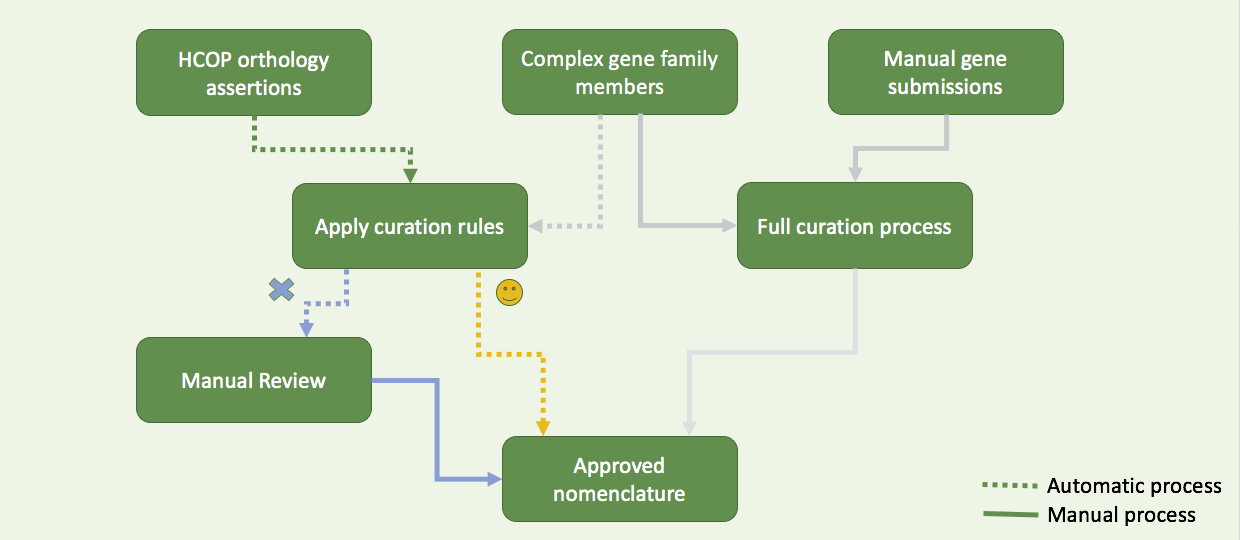
Data pipeline
4 resources
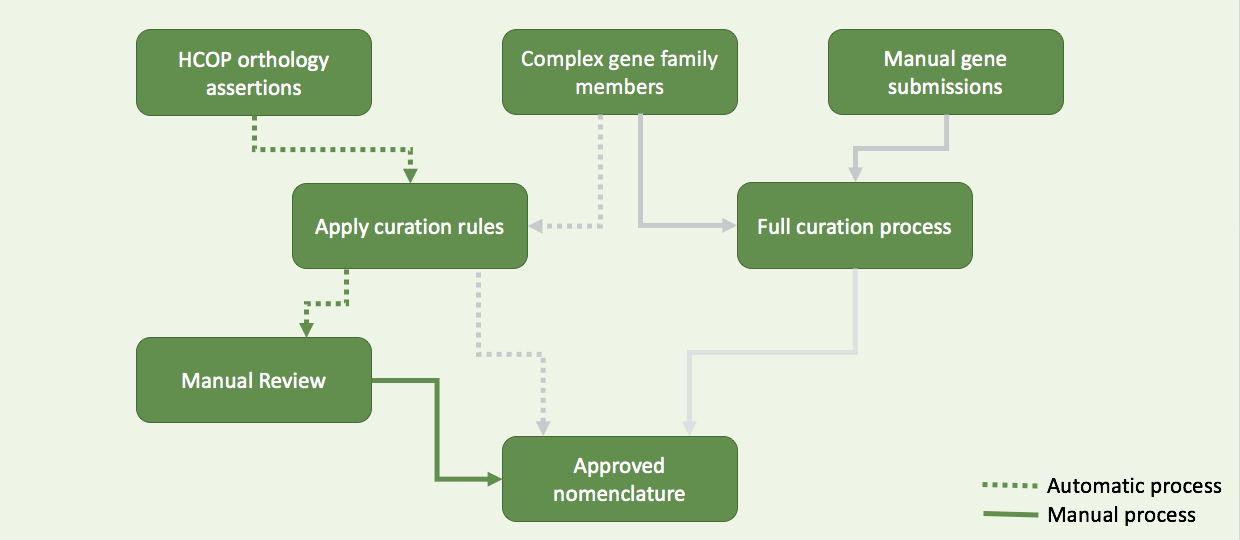
3 resources
Complex gene family
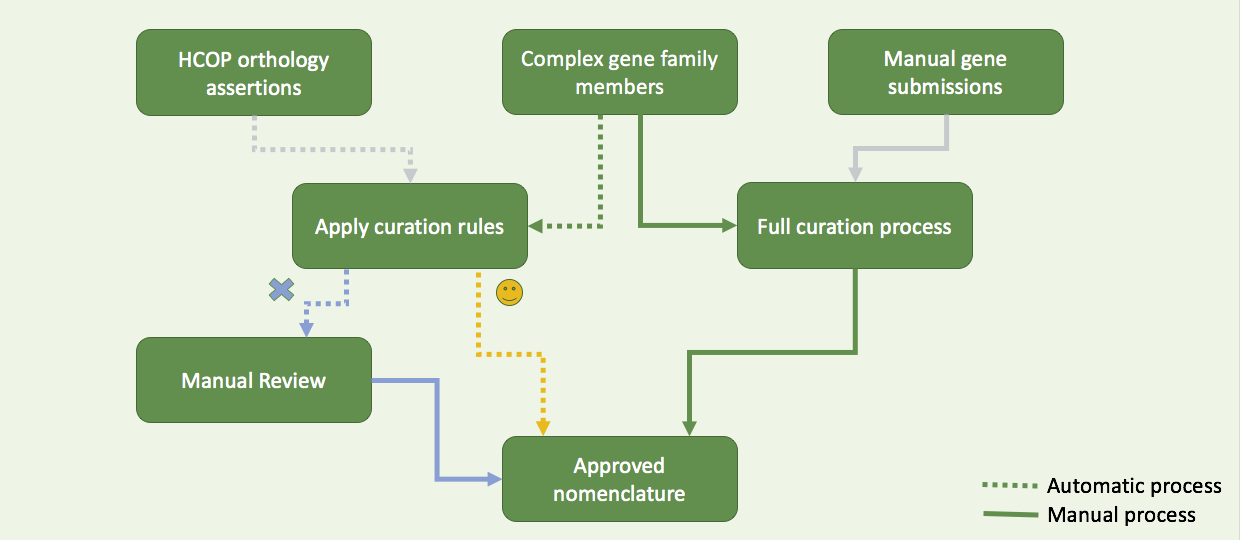
Manual submission
Prototype species: Chimp
- We have ~14800 named Chimp genes within the VGNC site.
- Most are 1:1 Human to Chimp orthologs with 4 resources agreeing
- Manually checked the results of the automation to test the pipeline.

Website
- Design is taken from the new HGNC site
- Branch of the same project to maximise code reuse
Future plans

HGNC curation
- Naming of novel loci that have been predicted to be protein coding by Gencode.
- Name lncRNAs if the models from Gencode and RefSeq are similar e.g LINC#.
- Continue to name small RNAs in collaboration the resources.
- Work in collaboration with TGMI to stabilise as many clinically relevant protein coding genes.
- Work towards renaming all uninformative human symbols like C#orf#.
HGNC informatics
- Release the new site.
- Make all the site HTTPS.
- Update Angular from 1.6 to 2+.
- Replace the old backend postgresql DB with a newly designed MySQL DB.
- Expand the REST API to include gene families.
- Possibly replace BioMart.
- Refactor the update pipeline.
VGNC
Continue curating nomenclature of remaining Chimp genes.
Add new species data
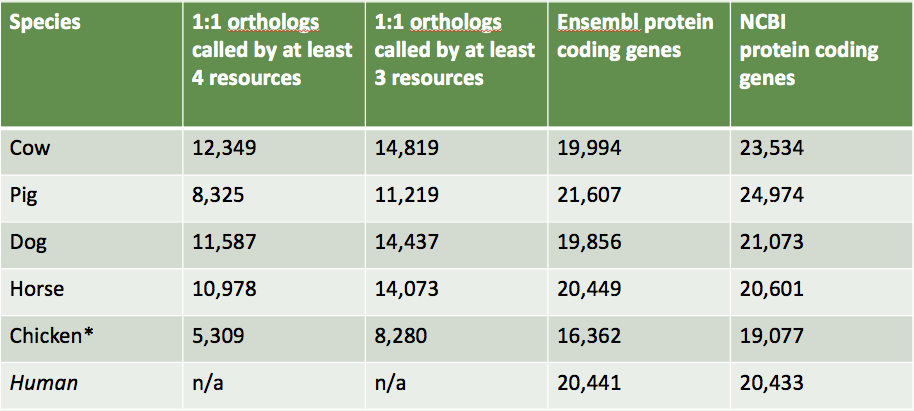
~10000 protein coding genes named automatically.
Species to work on next:
Dog

Cow

Horse

Expand on curation tools
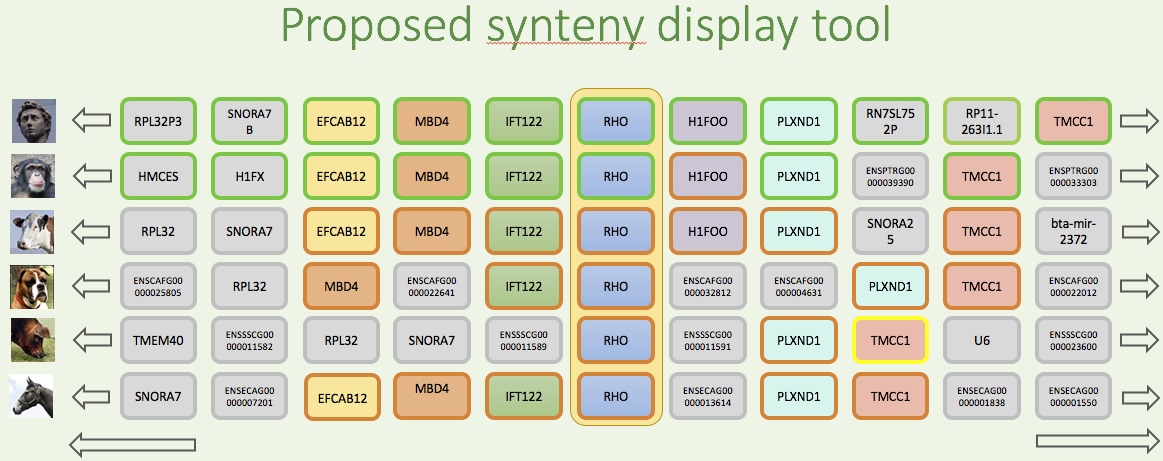
REST API
Need to create a REST API similar to the HGNC.
Utilise machine learning
- Investigate possible use of neural networks or SVMs.
- To improve the VGNC pipeline.
- For naming RNAs and complex families.

Thanks for listening

